| Attachment | Size |
|---|---|
| 2.06 MB |
Work by the authors described in this review was funded by grants from the Belgian FNRS, the European Research Council (ERC Adv Grant Gendevocortex), the WELBIO Program of the Walloon Region, the AXA Research Fund, the ERA-net 'Microkin', and the Vlaams Instituut voor Biotechnologie (VIB) (to P.V.). I.K.S. was supported by EMBO and FNRS Fellowships.
Brunetti-Pierri, N., Berg, J.S., Scaglia, F., Belmont, J., Bacino, C.A., Sahoo, T., Lalani, S.R., Graham, B., Lee, B., Shinawi, M., et al. (2008). Recurrent reciprocal 1q21.1 deletions and duplications associated with microcephaly or macrocephaly and developmental and behavioral abnormalities. Nat Genet 40, 1466-1471.
Cardone, M.F., Jiang, Z., D'Addabbo, P., Archidiacono, N., Rocchi, M., Eichler, E.E., and Ventura, M. (2008). Hominoid chromosomal rearrangements on 17q map to complex regions of segmental duplication. Genome Biol 9, R28.
Carroll, S.B. (2003). Genetics and the making of Homo sapiens. Nature 422, 849-857.
Carroll, S.B. (2008). Evo-devo and an expanding evolutionary synthesis: a genetic theory of morphological evolution. Cell 134, 25-36.
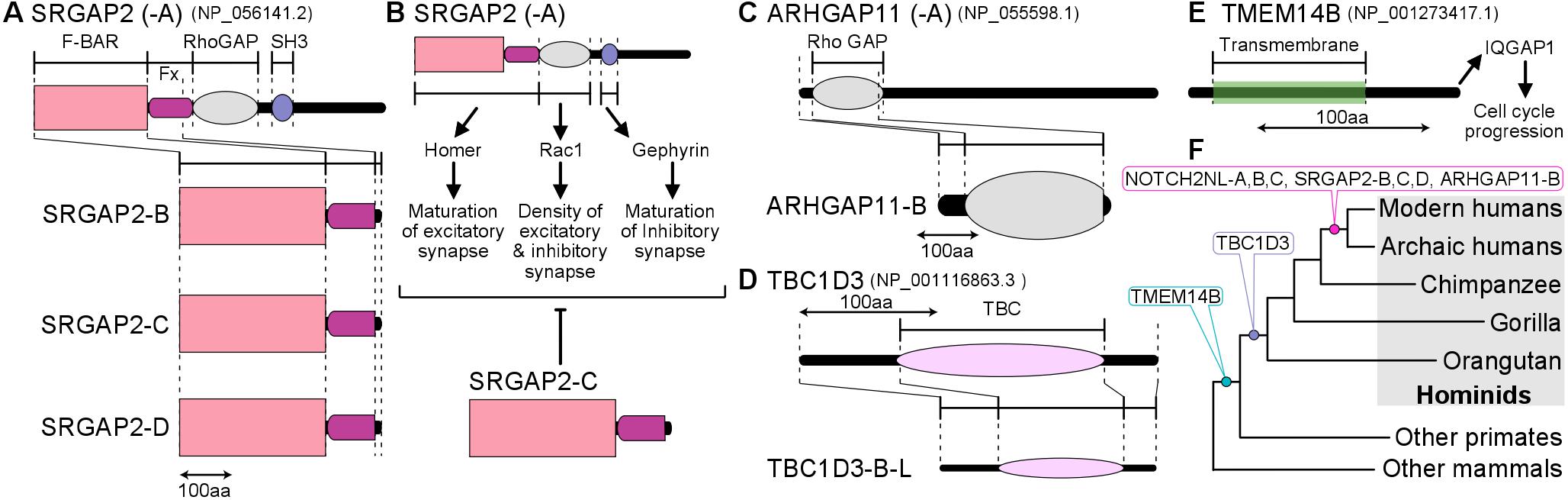
Charrier, C., Joshi, K., Coutinho-Budd, J., Kim, J.E., Lambert, N., de Marchena, J., Jin, W.L., Vanderhaeghen, P., Ghosh, A., Sassa, T., et al. (2012). Inhibition of SRGAP2 Function by Its Human-Specific Paralogs Induces Neoteny during Spine Maturation. Cell 149, 923-935.
Coe, B.P., Girirajan, S., and Eichler, E.E. (2012). The genetic variability and commonality of neurodevelopmental disease. Am J Med Genet C Semin Med Genet 160C, 118-129.
de la Torre-Ubieta, L., Stein, J.L., Won, H., Opland, C.K., Liang, D., Lu, D., and Geschwind, D.H. (2018). The Dynamic Landscape of Open Chromatin during Human Cortical Neurogenesis. Cell 172, 289-304 e218.
Defelipe, J. (2011). The evolution of the brain, the human nature of cortical circuits, and intellectual creativity. Frontiers in neuroanatomy 5, 29.
Dennis, M.Y., and Eichler, E.E. (2016). Human adaptation and evolution by segmental duplication. Current opinion in genetics & development 41, 44-52.
Dennis, M.Y., Harshman, L., Nelson, B.J., Penn, O., Cantsilieris, S., Huddleston, J., Antonacci, F., Penewit, K., Denman, L., Raja, A., et al. (2017). The evolution and population diversity of human-specific segmental duplications. Nat Ecol Evol 1, 69.
Dennis, M.Y., Nuttle, X., Sudmant, P.H., Antonacci, F., Graves, T.A., Nefedov, M., Rosenfeld, J.A., Sajjadian, S., Malig, M., Kotkiewicz, H., et al. (2012). Evolution of human-specific neural SRGAP2 genes by incomplete segmental duplication. Cell 149, 912-922.
Dougherty, M.L., Nuttle, X., Penn, O., Nelson, B.J., Huddleston, J., Baker, C., Harshman, L., Duyzend, M.H., Ventura, M., Antonacci, F., et al. (2017). The birth of a human-specific neural gene by incomplete duplication and gene fusion. Genome Biol 18, 49.
Dougherty, M.L., Underwood, J.G., Nelson, B.J., Tseng, E., Munson, K.M., Penn, O., Nowakowski, T.J., Pollen, A.A., and Eichler, E.E. (2018). Transcriptional fates of human-specific segmental duplications in brain. Genome Res 28, 1566-1576.
Du, A., Zipkin, A.M., Hatala, K.G., Renner, E., Baker, J.L., Bianchi, S., Bernal, K.H., and Wood, B.A. (2018). Pattern and process in hominin brain size evolution are scale-dependent. Proc R Soc B 285, 20172738.
Duan, Z., Li, F.-Q., Wechsler, J., Meade-White, K., Williams, K., Benson, K.F., and Horwitz, M. (2004). A novel notch protein, N2N, targeted by neutrophil elastase and implicated in hereditary neutropenia. Molecular and cellular biology 24, 58-70.
Dumas, L.J., O'Bleness, M.S., Davis, J.M., Dickens, C.M., Anderson, N., Keeney, J.G., Jackson, J., Sikela, M., Raznahan, A., Giedd, J., et al. (2012). DUF1220-domain copy number implicated in human brain-size pathology and evolution. American journal of human genetics 91, 444-454.
Espuny-camacho, I., Michelsen, K.A., Gall, D., Linaro, D., Hasche, A., Lambert, N., Gaspard, N., Pe, S., Bali, C., Orduz, D., et al. (2013). Pyramidal neurons derived from human pluripotent stem cells integrate efficiently into mouse brain circuits in vivo. Neuron 77, 440-456.
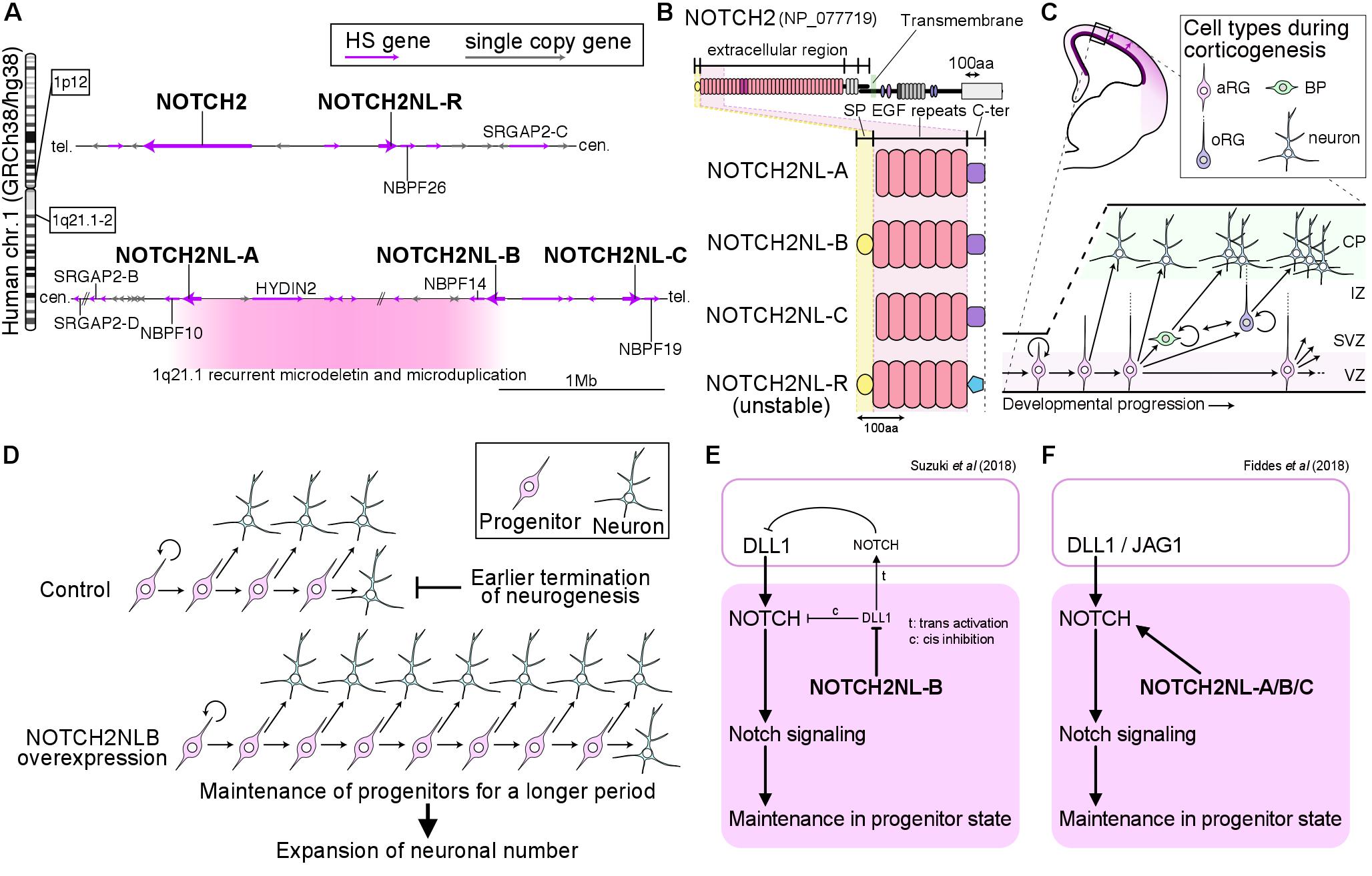
Fiddes, I.T., Lodewijk, G.A., Mooring, M.M., Bosworth, C.M., Ewing, A.D., Mantalas, G.L., Novak, A.M., van den Bout, A., Bishara, A., Rosenkrantz, J.L., et al. (2018). Human-specific NOTCH-like genes in a region linked to neurodevelopmental disorders affect cortical neurogenesis. Cell 173, 1356–1369.
Fietz, S.A., Lachmann, R., Brandl, H., Kircher, M., Samusik, N., Schröder, R., Lakshmanaperumal, N., Henry, I., Vogt, J., and Riehn, A. (2012). Transcriptomes of germinal zones of human and mouse fetal neocortex suggest a role of extracellular matrix in progenitor self-renewal. Proceedings of the National Academy of Sciences 109, 11836-11841.
Florio, M., Albert, M., Taverna, E., Namba, T., Brandl, H., Lewitus, E., Haffner, C., Sykes, A., Wong, F.K., Peters, J., et al. (2015). Human-specific gene ARHGAP11B promotes basal progenitor amplification and neocortex expansion. Science 347, 1465-1470.
Florio, M., Borrell, V., and Huttner, W.B. (2017). Human-specific genomic signatures of neocortical expansion. Current opinion in neurobiology 42, 33-44.
Florio, M., Heide, M., Pinson, A., Brandl, H., Albert, M., Winkler, S., Wimberger, P., Huttner, W.B., and Hiller, M. (2018). Evolution and cell-type specificity of human-specific genes preferentially expressed in progenitors of fetal neocortex. eLife 7, e32332.
Florio, M., Namba, T., Paabo, S., Hiller, M., and Huttner, W.B. (2016). A single splice site mutation in human-specific ARHGAP11B causes basal progenitor amplification. Science advances 2, e1601941.
Fossati, M., Pizzarelli, R., Schmidt, E.R., Kupferman, J.V., Stroebel, D., Polleux, F., and Charrier, C. (2016). SRGAP2 and Its Human-Specific Paralog Co-Regulate the Development of Excitatory and Inhibitory Synapses. Neuron 91, 356-369.
Frittoli, E., Palamidessi, A., Pizzigoni, A., Lanzetti, L., Garre, M., Troglio, F., Troilo, A., Fukuda, M., Di Fiore, P.P., Scita, G., et al. (2008). The primate-specific protein TBC1D3 is required for optimal macropinocytosis in a novel ARF6-dependent pathway. Molecular biology of the cell 19, 1304-1316.
Grandbarbe, L., Bouissac, J., Rand, M., de Angelis, M.H., Artavanis-Tsakonas, S., and Mohier, E. (2003). Delta-Notch signaling controls the generation of neurons/glia from neural stem cells in a stepwise process. Development 130, 1391-1402.
Grayton, H.M., Fernandes, C., Rujescu, D., and Collier, D.A. (2012). Copy number variations in neurodevelopmental disorders. Prog Neurobiol 99, 81-91.
Guerrier, S., Coutinho-Budd, J., Sassa, T., Gresset, A., Jordan, N.V., Chen, K., Jin, W.L., Frost, A., and Polleux, F. (2009). The F-BAR domain of srGAP2 induces membrane protrusions required for neuronal migration and morphogenesis. Cell 138, 990-1004.
Hatakeyama, J., Wakamatsu, Y., Nagafuchi, A., Kageyama, R., Shigemoto, R., and Shimamura, K. (2014). Cadherin-based adhesions in the apical endfoot are required for active Notch signaling to control neurogenesis in vertebrates. Development 141, 1671-1682.
Hill, R.S., and Walsh, C.A. (2005). Molecular insights into human brain evolution. Nature 437, 64-67.
Hodzic, D., Kong, C., Wainszelbaum, M.J., Charron, A.J., Su, X., and Stahl, P.D. (2006). TBC1D3, a hominoid oncoprotein, is encoded by a cluster of paralogues located on chromosome 17q12. Genomics 88, 731-736.
Johnson, M.B., Kawasawa, Y.I., Mason, C.E., Krsnik, Z., Coppola, G., Bogdanovic, D., Geschwind, D.H., Mane, S.M., State, M.W., and Sestan, N. (2009). Functional and evolutionary insights into human brain development through global transcriptome analysis. Neuron 62, 494-509.
Ju, X.C., Hou, Q.Q., Sheng, A.L., Wu, K.Y., Zhou, Y., Jin, Y., Wen, T., Yang, Z., Wang, X., and Luo, Z.G. (2016). The hominoid-specific gene TBC1D3 promotes generation of basal neural progenitors and induces cortical folding in mice. eLife 5.
Kageyama, R., Ohtsuka, T., Shimojo, H., and Imayoshi, I. (2008). Dynamic Notch signaling in neural progenitor cells and a revised view of lateral inhibition. Nat Neurosci 11, 1247-1251.
Kaminsky, E.B., Kaul, V., Paschall, J., Church, D.M., Bunke, B., Kunig, D., Moreno-De-Luca, D., Moreno-De-Luca, A., Mulle, J.G., Warren, S.T., et al. (2011). An evidence-based approach to establish the functional and clinical significance of copy number variants in intellectual and developmental disabilities. Genetics in medicine : official journal of the American College of Medical Genetics 13, 777-784.
Kawaguchi, D., Yoshimatsu, T., Hozumi, K., and Gotoh, Y. (2008). Selection of differentiating cells by different levels of delta-like 1 among neural precursor cells in the developing mouse telencephalon. Development 135, 3849-3858.
Khaitovich, P., Hellmann, I., Enard, W., Nowick, K., Leinweber, M., Franz, H., Weiss, G., Lachmann, M., and Pääbo, S. (2005). Parallel patterns of evolution in the genomes and transcriptomes of humans and chimpanzees. Science 309, 1850-1854.
Kronenberg, Z.N., Fiddes, I.T., Gordon, D., Murali, S., Cantsilieris, S., Meyerson, O.S., Underwood, J.G., Nelson, B.J., Chaisson, M.J.P., Dougherty, M.L., et al. (2018). High-resolution comparative analysis of great ape genomes. Science 360.
Lambert, N., Lambot, M.A., Bilheu, A., Albert, V., Englert, Y., Libert, F., Noel, J.C., Sotiriou, C., Holloway, A.K., Pollard, K.S., et al. (2011). Genes expressed in specific areas of the human fetal cerebral cortex display distinct patterns of evolution. PLoS One 6, e17753.
Lindsell, C.E., Boulter, J., diSibio, G., Gossler, A., and Weinmaster, G. (1996). Expression Patterns ofJagged, Delta1, Notch1, Notch2,andNotch3Genes Identify Ligand–Receptor Pairs That May Function in Neural Development. Molecular and Cellular Neuroscience 8, 14-27.
Liu, J., Liu, W., Yang, L., Wu, Q., Zhang, H., Fang, A., Li, L., Xu, X., Sun, L., Zhang, J., et al. (2017). The Primate-Specific Gene TMEM14B Marks Outer Radial Glia Cells and Promotes Cortical Expansion and Folding. Cell stem cell 21, 635-649 e638.
Lui, J.H., Hansen, D.V., and Kriegstein, A.R. (2011). Development and evolution of the human neocortex. Cell 146, 18-36.
Malhotra, D., and Sebat, J. (2012). CNVs: harbingers of a rare variant revolution in psychiatric genetics. Cell 148, 1223-1241.
McLean, C.Y., Reno, P.L., Pollen, A.A., Bassan, A.I., Capellini, T.D., Guenther, C., Indjeian, V.B., Lim, X., Menke, D.B., Schaar, B.T., et al. (2011). Human-specific loss of regulatory DNA and the evolution of human-specific traits. Nature 471, 216-219.
Mefford, H.C., and Eichler, E.E. (2009). Duplication hotspots, rare genomic disorders, and common disease. Curr Opin Genet Dev 19, 196-204.
Mefford, H.C., Sharp, A.J., Baker, C., Itsara, A., Jiang, Z., Buysse, K., Huang, S., Maloney, V.K., Crolla, J.A., and Baralle, D. (2008). Recurrent rearrangements of chromosome 1q21. 1 and variable pediatric phenotypes. New England Journal of Medicine 359, 1685-1699.
Miller, A.C., Lyons, E.L., and Herman, T.G. (2009). cis-Inhibition of Notch by endogenous Delta biases the outcome of lateral inhibition. Current Biology 19, 1378-1383.
Mora-Bermúdez, F., Badsha, F., Kanton, S., Camp, J.G., Vernot, B., Köhler, K., Voigt, B., Okita, K., Maricic, T., and He, Z. (2016). Differences and similarities between human and chimpanzee neural progenitors during cerebral cortex development. eLife 5.
Nelson, B.R., Hodge, R.D., Bedogni, F., and Hevner, R.F. (2013). Dynamic interactions between intermediate neurogenic progenitors and radial glia in embryonic mouse neocortex: potential role in Dll1-Notch signaling. Journal of Neuroscience 33, 9122-9139.
Nord, A.S., Pattabiraman, K., Visel, A., and Rubenstein, J.L.R. (2015). Genomic perspectives of transcriptional regulation in forebrain development. Neuron 85, 27-47.
O'Bleness, M., Searles, V.B., Varki, A., Gagneux, P., and Sikela, J.M. (2012). Evolution of genetic and genomic features unique to the human lineage. Nat Rev Genet 13, 853-866.
Ohno, S. (1970). Evolution by gene duplication (Springer).
Ohno, S. (1999). Gene duplication and the uniqueness of vertebrate genomes circa 1970-1999. Semin Cell Dev Biol 10, 517-522.
Pei, L., Peng, Y., Yang, Y., Ling, X.B., van Eyndhoven, W.G., Nguyen, K.C.Q., Rubin, M., Hoey, T., Powers, S., and Li, J. (2002). PRC17, a Novel Oncogene Encoding a Rab GTPase-activating Protein, Is Amplified in Prostate Cancer. Cancer research 62, 5420 –5424.
Petanjek, Z., Judas, M., Simic, G., Rasin, M.R., Uylings, H.B., Rakic, P., and Kostovic, I. (2011). Extraordinary neoteny of synaptic spines in the human prefrontal cortex. Proceedings of the National Academy of Sciences of the United States of America 108, 13281-13286.
Rakic, P. (2009). Evolution of the neocortex: a perspective from developmental biology. Nat Rev Neurosci 10, 724-735.
Sharp, A.J., Hansen, S., Selzer, R.R., Cheng, Z., Regan, R., Hurst, J.A., Stewart, H., Price, S.M., Blair, E., Hennekam, R.C., et al. (2006). Discovery of previously unidentified genomic disorders from the duplication architecture of the human genome. Nat Genet 38, 1038-1042.
Shubin, N., Tabin, C., and Carroll, S. (2009). Deep homology and the origins of evolutionary novelty. Nature 457, 818-823.
Sporny, M., Guez-Haddad, J., Kreusch, A., Shakartzi, S., Neznansky, A., Cross, A., Isupov, M.N., Qualmann, B., Kessels, M.M., and Opatowsky, Y. (2017). Structural History of Human SRGAP2 Proteins. Molecular biology and evolution 34, 1463-1478.
Sprinzak, D., Lakhanpal, A., LeBon, L., Santat, L.A., Fontes, M.E., Anderson, G.A., Garcia-Ojalvo, J., and Elowitz, M.B. (2010). Cis-interactions between Notch and Delta generate mutually exclusive signalling states. Nature 465, 86-90.
Stahl, P.D., and Wainszelbaum, M.J. (2009). Human-specific genes may offer a unique window into human cell signaling. Science signaling 2, pe59.
Stankiewicz, P., and Lupski, J.R. (2010). Structural variation in the human genome and its role in disease. Annual review of medicine 61, 437-455.
Steinberg, K.M., Schneider, V.A., Graves-Lindsay, T.A., Fulton, R.S., Agarwala, R., Huddleston, J., Shiryev, S.A., Morgulis, A., Surti, U., Warren, W.C., et al. (2014). Single haplotype assembly of the human genome from a hydatidiform mole. Genome Res 24, 2066-2076.
Sudmant, P.H., Kitzman, J.O., Antonacci, F., Alkan, C., Malig, M., Tsalenko, A., Sampas, N., Bruhn, L., Shendure, J., Genomes, P., et al. (2010). Diversity of human copy number variation and multicopy genes. Science 330, 641-646.
Suzuki, I.K., Gacquer, D., Van Heurck, R., Kumar, D., Wojno, M., Bilheu, A., Herpoel, A., Lambert, N., Cheron, J., Polleux, F., et al. (2018). Human-specific NOTCH2NL genes expand cortical neurogenesis through Delta/Notch regulation. Cell 173, 1370-1384. e1316.
Suzuki, I.K., and Vanderhaeghen, P. (2015). Is this a brain which I see before me? Modeling human neural development with pluripotent stem cells. Development 142, 3138.
Takumi, T., and Tamada, K. (2018). CNV biology in neurodevelopmental disorders. Curr Opin Neurobiol 48, 183-192.
Wainszelbaum, M.J., Charron, A.J., Kong, C., Kirkpatrick, D.S., Srikanth, P., Barbieri, M.A., Gygi, S.P., and Stahl, P.D. (2008). The hominoid-specific oncogene TBC1D3 activates Ras and modulates epidermal growth factor receptor signaling and trafficking. The Journal of biological chemistry 283, 13233-13242.
Weischenfeldt, J., Symmons, O., Spitz, F., and Korbel, J.O. (2013). Phenotypic impact of genomic structural variation: insights from and for human disease. Nat Rev Genet 14, 125-138.
Wilsch-Brauninger, M., Florio, M., and Huttner, W.B. (2016). Neocortex expansion in development and evolution - from cell biology to single genes. Curr Opin Neurobiol 39, 122-132.
Yoon, K., and Gaiano, N. (2005). Notch signaling in the mammalian central nervous system: insights from mouse mutants. Nat Neurosci 8, 709-715.
Yun, K., Fischman, S., Johnson, J., de Angelis, M.H., Weinmaster, G., and Rubenstein, J.L. (2002). Modulation of the notch signaling by Mash1 and Dlx1/2 regulates sequential specification and differentiation of progenitor cell types in the subcortical telencephalon. Development 129, 5029-5040.