The rapid advance of super-resolution microscopy and its experimental applications has provided neuroscientists with a pass to the nanoscopic world of synaptic machinery. Here we will briefly overview and discuss current progress in our understanding of the three-dimensional synaptic architecture and molecular organisation as gleaned from the imaging methods that go beyond the diffraction limit of conventional light microscopy. We will argue that such methods are to take our knowledge of synapses to a qualitatively new level, providing the neuroscience research community with novel organising principles and concepts pertinent to the workings of the brain.
Introduction
More than one hundred years ago, Santiago Ramón y Cajal used improved silver staining to image, for the first time, individual nerve cells, thus prompting the notion of synapses that separate them physically while connecting them functionally. Nevertheless, only the advance of electron microscopy (EM) in the 1950s made it possible to study the intricate details of pre- and postsynaptic structures that separate the two communicating neurons (Harris and Weinberg 2012; Palay and Palade 1955). However, EM can only be performed on fixed tissue and thus can be used to report dynamic processes only correlatively, by comparing different preparations.
The advance of fluorescence recovery after photobleaching (FRAP) and single-particle tracking (SPT) techniques paved the way for the study of dynamic changes in microscopic cellular compartments at super-resolution (SR), i.e. beyond the diffraction limit of conventional optical microscopy. In brief, the FRAP method can reveal the kinetics of diffusion of the (fluorescently-labelled) molecule of interest as well as the proportion of mobile and immobile fractions of the molecule species in the area (Axelrod et al. 1976). FRAP has been used to monitor the molecular kinetics and the morphology of both pre- and postsynaptic compartments (Henkel et al. 1996; Perestenko and Henley 2006; Rusakov et al. 2011). In turn, SPT methods rely on the labelling of individual molecules with bright fluorescent nanoparticles, such as quantum dots, in live cells (Groc et al. 2007; Medintz et al. 2005). SPT techniques revealed much detail about the molecular nature of excitatory as well as inhibitory synapses and their neural receptors (Choquet and Triller 2013; Hosy et al. 2014; Salvatico et al. 2015). By registering rare intermittent “blinking” (random switching) of individual nanoparticles the SPT method achieves nanoscale localisation and tracking of individual molecules for prolonged periods of time. Even though most SPT work has been done in neuronal culture preparations, it was recently adapted for organotypic (Biermann et al. 2014) as well as for acute brain slices and in vivo application (Varela et al. 2016). The implementation of other recently developed SR microscopy techniques has enabled localisation of individual molecules at synapses (Dani et al. 2010; Maglione and Sigrist 2013; Maidorn et al. 2016; Sigal et al. 2015; Tang et al. 2016; Willig and Barrantes 2014; Zhong 2015), at resolution as high as a few dozen nanometres (Minoshima and Kikuchi 2017; Sydor et al. 2015; Turkowyd et al. 2016; Yamanaka et al. 2014). Here we will provide an overview of some notable findings made by visualising synaptic structures with SR microscopy. We will focus on work in mammalian cells and tissue. However, some great work has been done in the model organism Drosophila, mainly imaging the molecular architecture of the neuromuscular junction (NMJ) (Bohme et al. 2016; Ehmann et al. 2014; Liu et al. 2011).
Below are some brief methodological details pertinent to these emerging techniques: for further insight into the physical principles of the SR techniques we refer the reader to the excellent reviews listed above. Stimulated-emission depletion (STED) microscopy uses doughnut-shaped focal illumination to partly deplete the excitation at its periphery. This narrows the emission spots thus enhancing resolution much beyond the diffraction limit (Klar et al. 2000). Another group of SR techniques is collectively called single molecule localisation microscopy (SMLM) and comprises two generic approaches (and their specific applications), photo-activated localization microscopy (PALM) and stochastic optical reconstruction microscopy (STORM). These methods are based on localising the point source of fluorescence (which can be one molecule) from its detected image (Betzig et al. 2006; Folling et al. 2008; Rust et al. 2006). Repeating stochastic excitation of a small and sparsely distributed fraction of fluorescent molecules per imaging cycle provides non-overlapping point sources of fluorescence and thus allows reconstruction of the exact position of the fluorescent molecules. These techniques can not only be used in fixed tissue but also in living cells, to study the real-time dynamics of individual molecules (Bethge et al. 2013; Jones et al. 2011; Owen et al. 2012). Moreover, one type of SMLM (sptPALM) uses the overexpression of a photoactivatable fluorescent protein fused to the protein of interest: this enables the imaging of hundreds of mini-trajectories for several milliseconds (Manley et al. 2008; Rossier et al. 2012). Another method, universal point accumulation for imaging in nanoscale topography (uPAINT), relies on the addition of fluorophores during the imaging process, which provides the dynamic imaging of continuously labelled, arbitrary membrane biomolecules in living cells (Giannone et al. 2010). In contrast to the nanoparticle-based SPT, which follows a few molecules for prolonged periods of time, sptPALM and uPAINT image single-molecule trajectories at very high densities for short periods of time (Sibarita 2014).
Collectively, SR imaging techniques allow the assessment of location, movement and counting the number of proteins of interest in living or fixed cells or tissue, thus deciphering the composition of protein complexes with nanometre spatial resolution and millisecond temporal resolution (Hosy et al. 2014; Minoshima and Kikuchi 2017; Sydor et al. 2015; Turkowyd et al. 2016; Yamanaka et al. 2014). This had not been possible with conventional imaging techniques. Several modifications have been made to the original SR protocols, with attention focusing on improving sample preparation, buffer mixtures and image analysis to adapt the techniques to image dynamic changes in living organisms. The present review will discuss several particular applications of such techniques focusing on the exploration of the presynaptic and the postsynaptic molecular organisation pertinent to the function and plasticity of central synapses.
Imaging of the vesicular release machinery using SR microscopy
The presynapse is a crowded yet dynamic compartment, in which the finely tuned regulation of molecular interactions is essential for its function in maintaining synaptic integrity. These processes include vesicular release and recycling of neurotransmitters; some aspects of which are still under debate (Maidorn et al. 2016; Rizzoli 2014). Here, single molecule imaging techniques should play an essential role in resolving some of these essential mechanisms.
Only a decade ago, detailed proteomic analyses using EM and mass spectrometry revealed the complexity of the synaptic vesicle proteome, generating a comprehensive list of the molecular players involved and a general model of their interactions (Sudhof 2006; Takamori et al. 2006). More recently, a comprehensive molecular model of an 'average' presynaptic axonal bouton was published, detailing the complexity of the system as a densely-packed gel (Wilhelm et al. 2014).
Several studies have focused on imaging the synaptic vesicles, their molecular composition and recycling mechanisms using different SR techniques (Kamin et al. 2010; Lehmann et al. 2015; Westphal et al. 2008; Willig et al. 2006). After synaptic vesicle fusion, some synaptotagmin molecules remained in the presynaptic membrane, which was interpreted as evidence for at least some synaptic vesicle constituents remaining together during recycling (Willig et al. 2006). Moreover, STED revealed that the plasma membrane SNAREs form clusters of ~60 nm diameter containing about 75 densely packed molecules in PC12 cells (Maglione and Sigrist 2013; Sieber et al. 2007). FRAP experiments detected freely diffusing molecules that dynamically move between such clusters (Sieber et al. 2007). Direct STORM (dSTORM) imaging showed that, indeed, the unclustered syntaxin as well as SNAP-25 molecules reside next to these clusters in PC12 cells (Bar-On et al. 2012). Furthermore, it has been shown that subsequent stimulation leads to endocytosis of a ‘readily retrievable’ pool of synaptic vesicles (Hua et al. 2011). Assemblies of synaptotagmin molecules in the plasmamembrane that might control membrane integrity or serve as molecular platforms for the formation of SNARE complexes have been identified in recent studies with STED (Maidorn et al. 2016; Milovanovic and Jahn 2015). STED was also used to visualise, for the first time, vesicular trafficking on the nanoscale in real time (Westphal et al. 2008). The buffer zone for endocytosis was also visualised in a study combining confocal and STED imaging, force measurements, pharmacology and gene knockout to investigate the dynamic assembly of filamentous actin that mediates Ω-profile vesicle merging at synapses (Wen et al. 2016). Additionally, actin dynamics were found to modulate presynaptic structure and function through dynamically controlling active zone organisation following local neuronal activity (Glebov et al. 2017). A recent study detected individual vesicle fusion events with ~27 nm localisation precision at single hippocampal synapses under physiological conditions (Maschi and Klyachko 2017). The researchers found that multiple distinct release sites could exist within individual synapses. These sites were distributed throughout the active zone and underwent repeated reuse in a dynamic manner that is activity dependent.
Besides vesicle dynamics, SR imaging was also employed to reveal the clustering of voltage-gated Ca2+ channels at synapses of auditory hair cells by bassoon (Frank et al. 2010) and to image the sandwich structure of piccolo and bassoon at NMJs in adult and aged mice (Nishimune et al. 2016) as well as the presence of brain-derived neurotrophic factor (BDNF) in small granules in the presynaptic face of excitatory synapses in cultured neurons (Andreska et al. 2014). A recent STED-based study investigated the presynaptic clustering of active zone proteins and revealed that RIM-BP2 is necessary for the clustering of calcium channels and bassoon at hippocampal CA3-CA1 synapses (Grauel et al. 2016). STORM imaging of hippocampal slices demonstrated that Δ9-tetrahydrocannabinol (THC) treatment induces internalisation and disappearance of type 1 cannabinoid receptor (CB1) receptors from gamma-aminobutyric acid (GABA)ergic axon terminals (Dudok et al. 2015). Younts et al. employed a combination of whole-cell recordings and STORM to investigate the role of local protein synthesis in rodent hippocampal slices. They found that presynaptic protein synthesis is required for long-term, but not short-term, plasticity of GABA release from CB1-expressing axons and might play a general role in controlling long-term plasticity in the mature mammalian brain (Younts et al. 2016). Moreover, a recent study found evidence for morphological plasticity of synaptic boutons and axon shafts to dynamically fine-tune action potential conduction velocity (Chereau et al. 2017).
Imaging the postsynaptic site of excitatory and inhibitory synapses using SR microscopy
Both STED and SMLM have been employed to analyse the structure of dendritic spines and the post-synapse. For example, in cultured neurons, STORM revealed the dynamics of thin filopodia and dendritic spines as well as of the endoplasmic reticulum and mitochondria (Shim et al. 2012). Additionally, STED has also been used to image the fine structure of dendritic spines (Berning et al. 2012; Bethge et al. 2013; Ding et al. 2009; Nagerl et al. 2008; Tonnesen et al. 2014; Tonnesen et al. 2011). In addition to revealing macroscopic changes of dendrites and spines, STED (D'Este et al. 2015; Sidenstein et al. 2016; Urban et al. 2011) and SMLM techniques (such as sptPALM) have been used to investigate actin filaments and their dynamics in spines and dendrites and their role in spine maturation and function (Chazeau et al. 2014; Frost et al. 2010; Izeddin et al. 2011; MacGillavry et al. 2016; Tatavarty et al. 2012; Tatavarty et al. 2009; Wang et al. 2016). It was found that dendritic actin polymerisation is more complex and heterogeneous than previously thought, with different actin flows and random motions (Frost et al. 2010; Tatavarty et al. 2009). Moreover, enhanced actin polymerisation was revealed in subdomains of the dendritic spine including the postsynaptic density (PSD) and the spine neck, which might play a role in the repositioning of glutamate receptors (Frost et al. 2010). Furthermore, the nucleation of actin extension seemed to start from a single point within the PSD, raising the possibility that changes in the environment are sensed by the actin filaments and in turn are translated into the PSD to mediate spine maturation (Chazeau et al. 2014; Izeddin et al. 2011). One of the organisers of mature spine structure is synaptopodin, which exerts an indirect effect, via F-actin, on the diffusion of membrane proteins in the spine neck (Wang et al. 2016). The lattice-like structures of filamentous actin were also imaged in vivo, revealing a structure similar to that seen in vitro (Berning et al. 2012; Willig et al. 2014). Altogether, SR techniques have substantially extended our view of the organisation of the spine cytoskeleton and revealed a much more crowded and organised cell compartment (Chazeau and Giannone 2016; MacGillavry and Hoogenraad 2015).
In addition to imaging the macroscopic structure of spines and the dynamics of the cytoskeleton within, SR techniques have helped to reveal the nanoscopic arrangement of the PSD. The first study investigating the nanoscale structure of excitatory synapses was performed in fixed rat brain tissue (Dani et al. 2010). The group studied the nanoscopic distribution of pre- and postsynaptic scaffolding proteins and established a detailed three-dimensional map of the synapses. Furthermore, they demonstrated variations in the organisation of N-methyl-D-aspartate receptors (NMDARs) and α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid receptors (AMPARs) across three different regions of the brain (Dani et al. 2010).
AMPARs are important for memory formation and synaptic plasticity, and their insertion into and removal from synapses is tightly regulated (MacGillavry et al. 2011; Newpher and Ehlers 2008). Their clustering in nanodomains at synapses was deciphered in 2013 by three independent groups using a variety of SR methods (Fukata et al. 2013; MacGillavry et al. 2013; Nair et al. 2013). All the groups have concluded that AMPARs are organized at synapses in nanoclusters much smaller than the PSD (0-4 AMPAR nanodomains/PSD with a homogenous size distribution of ~80 nm). The number of the AMPAR nanodomains depended on the size of the underlying PSD. The average centre-to-centre distance between the nanodomains is 500 nm so that a single glutamate release cannot act on more than one domain (Nair et al. 2013). The first of these publications used PALM of several postsynaptic scaffolding molecules such as PSD95 and Homer1 as well as dSTORM of AMPAR and NMDARs in live and fixed neuronal cultures (MacGillavry et al. 2013). The second paper used a combination of techniques in live cultures investigating the dynamics of palmytolated PSD95 (Fukata et al. 2013). The researchers used FRAP and PALM and found that the PSD was organized through subdomains of PSD95, which in turn are regulated through the DHHC2-dependent palmitoylation (Fukata et al. 2013). Lastly, one group used uPAINT, sptPALM, dSTORM and STED to discover that AMPARs are immobile at the aforementioned nanodomains but diffuse freely between them, which could be due to the nanoorganisation of PSD95 scaffolds (Nair et al. 2013). dSTORM data revealed that 20-25 AMPARs are located in each nanodomain. All groups also investigated the organisation of PSD95 molecules within the PSD. The results revealed that even though PSD95 can be found throughout the PSD, this protein, too, clusters in nanodomains of ~150 nm in diameter (Fukata et al. 2013; MacGillavry et al. 2013; Nair et al. 2013). These nanodomains of PSD95 were also revealed in brain slices (Broadhead et al. 2016; Tang et al. 2016). Using a single transmembrane domain with a PDZ binding motif, another group investigated the impact of the tight protein packing within the PSD and concluded that both crowding and binding dynamics limit diffusion and escape of AMPARs from the synapse (Li et al. 2016). Blocking the interaction of AMPARs with stargazin, which regulates AMPAR trafficking and gating, increased the rate of AMPAR diffusion as measured with sptPALM but revealed an interaction of the receptors with other proteins outside of the PSD (Hoze et al. 2012). Even though the diffusion properties of neural receptors had been investigated already in live cells using nanoparticle-based SPT the novel studies were the first to describe the nano-organisation of AMPARs and of scaffolding proteins at PSDs.
STED and SMLM have not only been applied to image the diffusion dynamics of AMPARs and PSD95 but also to assess the synaptic localisation and dynamics of other receptors, channels and scaffolding proteins. For example, it was found that the A-kinase-anchoring protein 79/150 clusters different ion channels (M-type K+, L-type Ca2+ and TRPV1) together with G protein-coupled receptors into functional domains as revealed by STORM (Zhang et al. 2016). STED revealed the size distribution, association into nanoclusters and long-range interactions of nicotinic acetyl-choline receptor nanoclusters and their cholesterol dependence (Kellner et al. 2007). The number and nano-organisation of the Na+K+-ATPase at excitatory synapses as well as clustering and co-localisation with dopamine D1 receptor and DARPP-32 was assessed using STED and PALM (Blom et al. 2016; Blom et al. 2012; Blom et al. 2011; Blom et al. 2013). Researchers found that also the P2X7 receptor is stabilised in nanoclusters of ~100 nm diameter and that its trafficking shows confinement in synaptic areas (Shrivastava et al. 2013). Moreover, STED and sptPALM were employed to study the surface dynamics of calcium channel subunits in cultured cells (Voigt et al. 2016). Using a novel monomeric streptavidin dSTORM approach Chamma et al. investigated the relative distribution of presynaptic Nrx1β and its postsynaptic binding partners Nlg1 and LRRTM2 in primary hippocampal neurons (Chamma et al. 2016a; Chamma et al. 2016b).
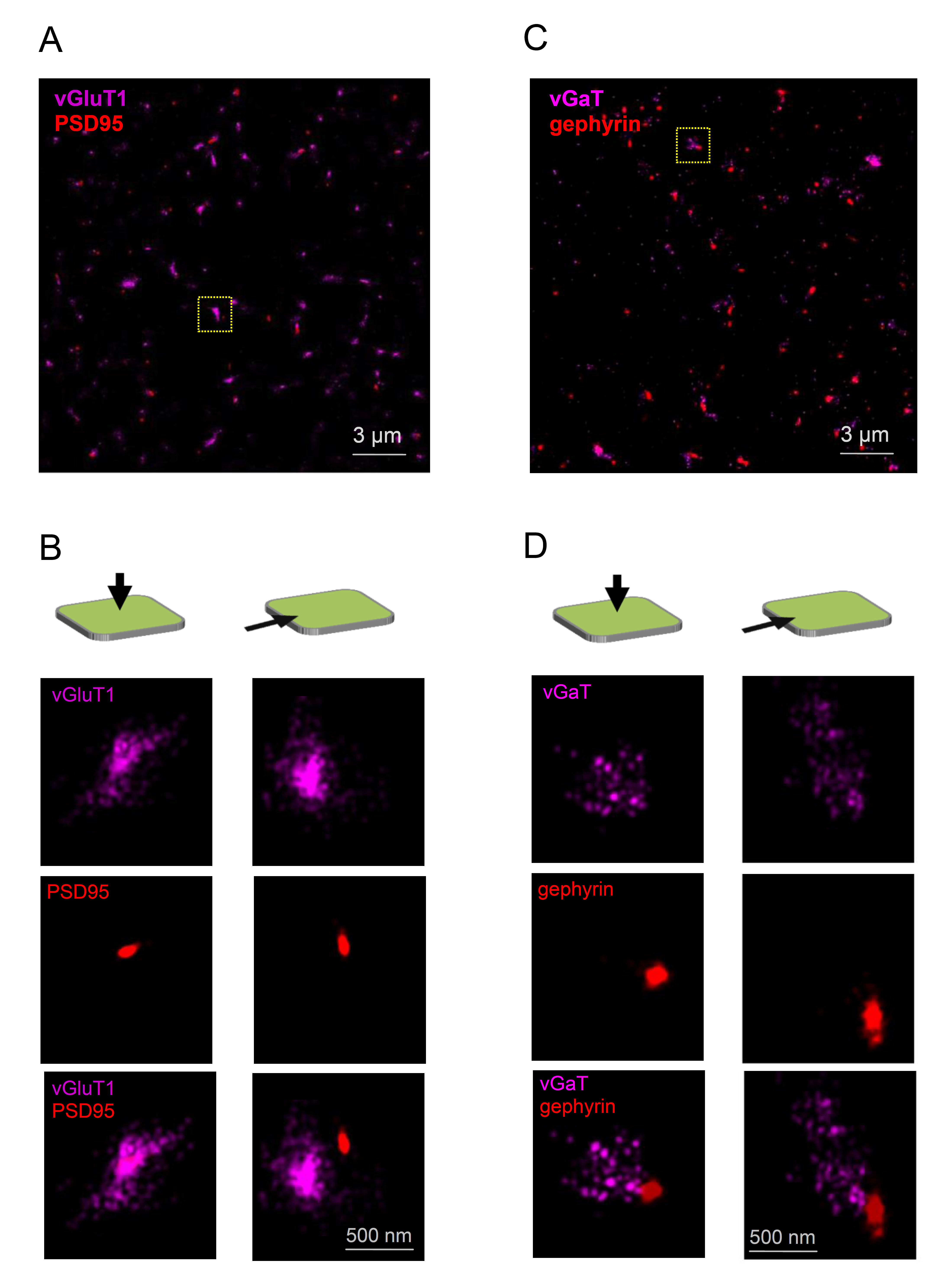
SR imaging has also been applied to visualise the sub-synaptic distribution of inhibitory synaptic components. For example, quantitative 3D-PALM analysis revealed that inhibitory synapses in cultured spinal cord neurons contain 40-500 gephyrin molecules, packed at a density of about 5000 molecules/µm2 (Specht et al. 2013). In native tissue, synapses contained about three times as many molecules and were packed more densely (Specht et al. 2013). A comparison of the nano-organisation of the postsynaptic scaffolding proteins gephyrin and PSD95 in cultured hippocampal neurons was performed using STED (Dzyubenko et al. 2016). An elegant study from the Zhuang lab combined STORM with array tomography to reconstruct the entire inhibitory synaptic input field of individual retinal ganglion and amacrine cells in mice and determined the spatial distribution and neurotransmitter receptor identity of the synapses therein (Sigal et al. 2015). An in-house example of a case study illustrating the 3D juxtaposition of two key pre- and postsynaptic proteins, both in excitatory and in inhibitory synapses, is shown in Figure 1. Whilst such approaches cannot achieve spatial resolution of EM, they could reveal the 3D distribution of individual protein molecules on a larger, contiguous scale, the task unattainable by EM techniques.
Figure 1. Nanoscopy of synaptic organisation in the hippocampus: a case study. (A) dSTORM images of 40 µm hippocampal sections in area CA1 (500 g male rat) imaged in a photoswitching buffer containing 100 mM cysteamine and oxygen scavengers (glucose oxidase and catalase) (Metcalf et al. 2013). Recorded with a Vutara 350 microscope (Bruker Corp., Billerica, US-MA) based on the single molecule localization (SML) biplane technology (Juette et al. 2008; Mlodzianoski et al. 2009). The microenvironment of individual synapses shown labelled with PSD95 (CF568-tagged, Novus NB300-556, red) and vesicular glutamate transporter vGluT1 (Alexa647-tagged, Merck Millipore AB5905, magenta) at a low magnification field of view. (B) One synaptic environment selected from the field of view (yellow dotted square in A) shown at higher magnification, at z-axis and x-axis projections, as indicated by arrows; separate and combined images of the two channels are shown. (C) dSTORM image depicting the microenvironment of inhibitory synapses labelled for gephyrin (CF568-tagged, SynapticSystems 147 021, red) and vGAT (Alexa647-tagged, SynapticSystems 131 004, magenta). Other conditions, settings and notations as in A. (D) One synaptic environment selected from the field of view (yellow dotted square in C) shown at higher magnification. Other notations as in B.
Combining synaptic plasticity and SR imaging
Recent reviews discussed the necessity of correlating electrophysiological recording at the single synapse level with SMLM to unveil the plastic pre- and postsynaptic organisation (Compans et al. 2016; Hosy et al. 2014). Several recent studies have assessed the dynamic changes of receptor clustering after synaptic stimulation. For example, during long-term potentiation of inhibitory transmission (iLTP), gephyrin is accumulated at inhibitory synapses and in turn leads to increased numbers of synaptic GABAA receptors (Petrini et al. 2014). This was further investigated by the same group using SR techniques (Pennacchietti et al. 2017). The researchers found that the nanoscale organisation of inhibitory synaptic proteins as an important determinant for inhibitory synaptic plasticity, thus adding a further level of complexity to the regulation of the neuronal network activity by plastic inhibitory synaptic signals (Pennacchietti et al. 2017). Moreover, Lu et al. used PALM and STORM to investigate the trafficking of CamKII in spines and revealed that the protein exists in three kinetic states: slow (interaction with immobile substrates), intermediate (binding to actin) and fast (CamKII alone). Stimulation of NMDARs slowed down these diffusion rates (Lu et al. 2014). Moreover, the phosphorylation of AMPARs by CamKII is crucial for P2X2-mediated AMPAR internalisation and ATP-driven synaptic depression (Pougnet et al. 2016; Pougnet et al. 2014). Moreover, Smith el al. found that ankyrin-G accumulates in spines after induction of chemical LTP and its knockdown prevented spine head enlargement (Smith et al. 2014). Chemical LTP also induces MMP-dependent enlargement of a subset of small spines and immobilisation, synaptic accumulation and clustering of AMPARs at spines (Szepesi et al. 2014). Furthermore, the cell adhesion molecule SynCAM 1 displayed increase in nanodomain size in long-term depression (Perez de Arce et al. 2015) and ephrin B3 was found to regulate the localisation of PSD95 to stable synapses and in stable nanodomains (Hruska et al. 2015). A recent publication revealed trans-synaptic nanocolumns that regulate the positioning of pre- and postsynaptic scaffolding proteins (Tang et al. 2016). The researchers found that the presynaptic protein RIM1/2 clusters in a very similar way to the postsynaptic nanoclusters that have been identified for PSD95 earlier (Broadhead et al. 2016; Fukata et al. 2013; MacGillavry et al. 2013; Nair et al. 2013) and that both nanodomains co-align across the synaptic cleft. Moreover, after LTP induction these nanocolumns remain with an increase in PSD95 clusters. This was the first study that provided proof for a trans-synaptic organisation of scaffolding molecules. However, the parameters and events that mediate this clustering remain still to be discovered. A recent elegant study combined PALM and optogenetics (Sinnen et al. 2017). Through optogenetically controlling the insertion of PSD95 molecules and hence AMPARs into established PSDs the researchers found that simple insertion of the receptors was not enough to drive potentiation of the synapses (Sinnen et al. 2017). Therefore, more complex changes in the nanodomain composition and characteristics are required for spine maturation and the gating of synaptic strength.
Concluding remarks
Several elegant studies have shed light on the nanoscopic events pertinent to synaptic plasticity, which were not possible to visualise without the advent of SR technology. However, most studies detailed here were conducted in cultured cells or fixed brain sections. Even though these results are undoubtedly very important the ultimate goal should be to achieve imaging in thick organised and living tissue. Some groups have already achieved this (Berning et al. 2012; Chen et al. 2014; Gao et al. 2012; Urban et al. 2011). Unfortunately, achieving SR in native tissue poses many difficulties such as restricted optical access, sample movement, or optical aberrations induced by the specimen. One way around these difficulties is adaptive optics (Booth et al. 2015), which improves the signal-to-noise ratio, axial resolution, the depth penetration as shown using in vivo two-photon excitation imaging (Ji et al. 2012; Rueckel et al. 2006; Wang et al. 2014) or STED (Gould et al. 2012; Patton et al. 2016). Further improvement to perform reliable multi-colour imaging will be important. Also, recreation of SPT work using SR in thicker tissue will be more relatable to physiology. 3D super-resolution microscopy can now effectively be employed in many laboratories using custom build or commercial microscopes; revealing structures way below the diffraction limit of light (Heller et al. 2017; Huang et al. 2008; Juette et al. 2008). The development and adaption of novel SR techniques such as qPAINT (Jungmann et al. 2016), integrating exchangeable single-molecule localization (IRIS) (Kiuchi et al. 2015), 4Pi single-molecule switching nanoscopy (Huang et al. 2016) and improvements such as the use of photoactivatable genetically encoded calcium indicators (Berlin et al. 2015), nanobodies (Pleiner et al. 2015), aptamers (de Castro et al. 2016), monomeric streptavidin for labelling (Chamma et al. 2017), or using the pore-forming bacterial toxin streptolysin O to label intracellular proteins in living mammalian cells for SR microscopy (Teng et al. 2016; Teng et al. 2017) will increase the resolution of these approaches even further to reveal even more intricate events. Also the recent invention of expansion microscopy (Chen et al. 2015; Chozinski et al. 2016; Tillberg et al. 2016) will make SR more affordable for many laboratories.
In future work, it will be important to elucidate the positions, clustering and general distribution patterns for the molecule of interest. The exciting new imaging techniques will help to reveal the nanoscale relationship in the tri-partite or tetra-partite synapse (Dityatev and Rusakov 2011). Recently, STED has been applied to image the tripartite synapse, focusing on plasticity of fine astrocytic processes (Panatier et al. 2014). Knowing the exact localisation and organisation of synapse-related molecules on neurons and glia cells will provide further information on the molecular mechanisms that govern plasticity of brain circuits, and may also offer novel insights into the development of new therapeutic strategies. SR imaging has already been applied to study disease models, such as Alzheimer’s (Schedin-Weiss et al. 2016; Siskova et al. 2014), intellectual disabilities (Barnes et al. 2015; Wijetunge et al. 2014), amyotrophic lateral sclerosis and frontotemporal dementia (Schoen et al. 2015). The new discoveries should help to translate these findings into the search of therapeutic intervention pertinent to human physiology.
| Attachment | Size |
|---|---|
| 878.89 KB |
This work was supported by the Wellcome Trust Principal Fellowship (101896), European Research Council Advanced Grant (323113-NETSIGNAL), FP7 ITN EXTRABRAIN Marie Curie Action (European Commission), Russian Science Foundation (15-14-30000), and Sumitomo Dainippon Pharma Co. Ltd. (Osaka, Japan).
Andreska T, Aufmkolk S, Sauer M, Blum R. 2014. High abundance of BDNF within glutamatergic presynapses of cultured hippocampal neurons. Front Cell Neurosci 8:107.
Axelrod D, Koppel DE, Schlessinger J, Elson E, Webb WW. 1976. Mobility measurement by analysis of fluorescence photobleaching recovery kinetics. Biophys J 16(9):1055-1069.
Bar-On D, Wolter S, van de Linde S, Heilemann M, Nudelman G, Nachliel E, Gutman M, Sauer M, Ashery U. 2012. Super-resolution imaging reveals the internal architecture of nano-sized syntaxin clusters. J Biol Chem 287(32):27158-27167.
Barnes SA, Wijetunge LS, Jackson AD, Katsanevaki D, Osterweil EK, Komiyama NH, Grant SG, Bear MF, Nagerl UV, Kind PC, Wyllie DJ. 2015. Convergence of Hippocampal Pathophysiology in Syngap+/- and Fmr1-/y Mice. J Neurosci 35(45):15073-15081.
Berlin S, Carroll EC, Newman ZL, Okada HO, Quinn CM, Kallman B, Rockwell NC, Martin SS, Lagarias JC, Isacoff EY. 2015. Photoactivatable genetically encoded calcium indicators for targeted neuronal imaging. Nat Methods 12(9):852-858.
Berning S, Willig KI, Steffens H, Dibaj P, Hell SW. 2012. Nanoscopy in a living mouse brain. Science 335(6068):551.
Bethge P, Chereau R, Avignone E, Marsicano G, Nagerl UV. 2013. Two-photon excitation STED microscopy in two colors in acute brain slices. Biophys J 104(4):778-785.
Betzig E, Patterson GH, Sougrat R, Lindwasser OW, Olenych S, Bonifacino JS, Davidson MW, Lippincott-Schwartz J, Hess HF. 2006. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313(5793):1642-1645.
Biermann B, Sokoll S, Klueva J, Missler M, Wiegert JS, Sibarita JB, Heine M. 2014. Imaging of molecular surface dynamics in brain slices using single-particle tracking. Nat Commun 5:3024.
Blom H, Bernhem K, Brismar H. 2016. Sodium pump organization in dendritic spines. Neurophotonics 3(4):041803.
Blom H, Ronnlund D, Scott L, Spicarova Z, Rantanen V, Widengren J, Aperia A, Brismar H. 2012. Nearest neighbor analysis of dopamine D1 receptors and Na(+)-K(+)-ATPases in dendritic spines dissected by STED microscopy. Microsc Res Tech 75(2):220-228.
Blom H, Ronnlund D, Scott L, Spicarova Z, Widengren J, Bondar A, Aperia A, Brismar H. 2011. Spatial distribution of Na+-K+-ATPase in dendritic spines dissected by nanoscale superresolution STED microscopy. BMC Neurosci 12:16.
Blom H, Ronnlund D, Scott L, Westin L, Widengren J, Aperia A, Brismar H. 2013. Spatial distribution of DARPP-32 in dendritic spines. PLoS One 8(9):e75155.
Bohme MA, Beis C, Reddy-Alla S, Reynolds E, Mampell MM, Grasskamp AT, Lutzkendorf J, Bergeron DD, Driller JH, Babikir H, Gottfert F, Robinson IM, O'Kane CJ, Hell SW, Wahl MC, Stelzl U, Loll B, Walter AM, Sigrist SJ. 2016. Active zone scaffolds differentially accumulate Unc13 isoforms to tune Ca(2+) channel-vesicle coupling. Nat Neurosci 19(10):1311-1320.
Booth M, Andrade D, Burke D, Patton B, Zurauskas M. 2015. Aberrations and adaptive optics in super-resolution microscopy. Microscopy (Oxf) 64(4):251-261.
Broadhead MJ, Horrocks MH, Zhu F, Muresan L, Benavides-Piccione R, DeFelipe J, Fricker D, Kopanitsa MV, Duncan RR, Klenerman D, Komiyama NH, Lee SF, Grant SG. 2016. PSD95 nanoclusters are postsynaptic building blocks in hippocampus circuits. Sci Rep 6:24626.
Chamma I, Letellier M, Butler C, Tessier B, Lim KH, Gauthereau I, Choquet D, Sibarita JB, Park S, Sainlos M, Thoumine O. 2016a. Mapping the dynamics and nanoscale organization of synaptic adhesion proteins using monomeric streptavidin. Nat Commun 7:10773.
Chamma I, Levet F, Sibarita JB, Sainlos M, Thoumine O. 2016b. Nanoscale organization of synaptic adhesion proteins revealed by single-molecule localization microscopy. Neurophotonics 3(4):041810.
Chamma I, Rossier O, Giannone G, Thoumine O, Sainlos M. 2017. Optimized labeling of membrane proteins for applications to super-resolution imaging in confined cellular environments using monomeric streptavidin. Nat Protoc 12(4):748-763.
Chazeau A, Giannone G. 2016. Organization and dynamics of the actin cytoskeleton during dendritic spine morphological remodeling. Cell Mol Life Sci 73(16):3053-3073.
Chazeau A, Mehidi A, Nair D, Gautier JJ, Leduc C, Chamma I, Kage F, Kechkar A, Thoumine O, Rottner K, Choquet D, Gautreau A, Sibarita JB, Giannone G. 2014. Nanoscale segregation of actin nucleation and elongation factors determines dendritic spine protrusion. EMBO J 33(23):2745-2764.
Chen BC, Legant WR, Wang K, Shao L, Milkie DE, Davidson MW, Janetopoulos C, Wu XS, Hammer JA, 3rd, Liu Z, English BP, Mimori-Kiyosue Y, Romero DP, Ritter AT, Lippincott-Schwartz J, Fritz-Laylin L, Mullins RD, Mitchell DM, Bembenek JN, Reymann AC, Bohme R, Grill SW, Wang JT, Seydoux G, Tulu US, Kiehart DP, Betzig E. 2014. Lattice light-sheet microscopy: imaging molecules to embryos at high spatiotemporal resolution. Science 346(6208):1257998.
Chen F, Tillberg PW, Boyden ES. 2015. Optical imaging. Expansion microscopy. Science 347(6221):543-548.
Chereau R, Saraceno GE, Angibaud J, Cattaert D, Nagerl UV. 2017. Superresolution imaging reveals activity-dependent plasticity of axon morphology linked to changes in action potential conduction velocity. Proc Natl Acad Sci U S A 114(6):1401-1406.
Choquet D, Triller A. 2013. The dynamic synapse. Neuron 80(3):691-703.
Chozinski TJ, Halpern AR, Okawa H, Kim HJ, Tremel GJ, Wong RO, Vaughan JC. 2016. Expansion microscopy with conventional antibodies and fluorescent proteins. Nat Methods 13(6):485-488.
Compans B, Choquet D, Hosy E. 2016. Review on the role of AMPA receptor nano-organization and dynamic in the properties of synaptic transmission. Neurophotonics 3(4):041811.
D'Este E, Kamin D, Gottfert F, El-Hady A, Hell SW. 2015. STED nanoscopy reveals the ubiquity of subcortical cytoskeleton periodicity in living neurons. Cell Rep 10(8):1246-1251.
Dani A, Huang B, Bergan J, Dulac C, Zhuang X. 2010. Superresolution imaging of chemical synapses in the brain. Neuron 68(5):843-856.
de Castro MA, Rammner B, Opazo F. 2016. Aptamer Stainings for Super-resolution Microscopy. Methods Mol Biol 1380:197-210.
Ding JB, Takasaki KT, Sabatini BL. 2009. Supraresolution imaging in brain slices using stimulated-emission depletion two-photon laser scanning microscopy. Neuron 63(4):429-437.
Dityatev A, Rusakov DA. 2011. Molecular signals of plasticity at the tetrapartite synapse. Curr Opin Neurobiol 21(2):353-359.
Dudok B, Barna L, Ledri M, Szabo SI, Szabadits E, Pinter B, Woodhams SG, Henstridge CM, Balla GY, Nyilas R, Varga C, Lee SH, Matolcsi M, Cervenak J, Kacskovics I, Watanabe M, Sagheddu C, Melis M, Pistis M, Soltesz I, Katona I. 2015. Cell-specific STORM super-resolution imaging reveals nanoscale organization of cannabinoid signaling. Nat Neurosci 18(1):75-86.
Dzyubenko E, Rozenberg A, Hermann DM, Faissner A. 2016. Colocalization of synapse marker proteins evaluated by STED-microscopy reveals patterns of neuronal synapse distribution in vitro. J Neurosci Methods 273:149-159.
Ehmann N, van de Linde S, Alon A, Ljaschenko D, Keung XZ, Holm T, Rings A, DiAntonio A, Hallermann S, Ashery U, Heckmann M, Sauer M, Kittel RJ. 2014. Quantitative super-resolution imaging of Bruchpilot distinguishes active zone states. Nat Commun 5:4650.
Folling J, Bossi M, Bock H, Medda R, Wurm CA, Hein B, Jakobs S, Eggeling C, Hell SW. 2008. Fluorescence nanoscopy by ground-state depletion and single-molecule return. Nat Methods 5(11):943-945.
Frank T, Rutherford MA, Strenzke N, Neef A, Pangrsic T, Khimich D, Fejtova A, Gundelfinger ED, Liberman MC, Harke B, Bryan KE, Lee A, Egner A, Riedel D, Moser T. 2010. Bassoon and the synaptic ribbon organize Ca2+ channels and vesicles to add release sites and promote refilling. Neuron 68(4):724-738.
Frost NA, Shroff H, Kong H, Betzig E, Blanpied TA. 2010. Single-molecule discrimination of discrete perisynaptic and distributed sites of actin filament assembly within dendritic spines. Neuron 67(1):86-99.
Fukata Y, Dimitrov A, Boncompain G, Vielemeyer O, Perez F, Fukata M. 2013. Local palmitoylation cycles define activity-regulated postsynaptic subdomains. J Cell Biol 202(1):145-161.
Gao L, Shao L, Higgins CD, Poulton JS, Peifer M, Davidson MW, Wu X, Goldstein B, Betzig E. 2012. Noninvasive imaging beyond the diffraction limit of 3D dynamics in thickly fluorescent specimens. Cell 151(6):1370-1385.
Giannone G, Hosy E, Levet F, Constals A, Schulze K, Sobolevsky AI, Rosconi MP, Gouaux E, Tampe R, Choquet D, Cognet L. 2010. Dynamic superresolution imaging of endogenous proteins on living cells at ultra-high density. Biophys J 99(4):1303-1310.
Glebov OO, Jackson RE, Winterflood CM, Owen DM, Barker EA, Doherty P, Ewers H, Burrone J. 2017. Nanoscale Structural Plasticity of the Active Zone Matrix Modulates Presynaptic Function. Cell Rep 18(11):2715-2728.
Gould TJ, Burke D, Bewersdorf J, Booth MJ. 2012. Adaptive optics enables 3D STED microscopy in aberrating specimens. Opt Express 20(19):20998-21009.
Grauel MK, Maglione M, Reddy-Alla S, Willmes CG, Brockmann MM, Trimbuch T, Rosenmund T, Pangalos M, Vardar G, Stumpf A, Walter AM, Rost BR, Eickholt BJ, Haucke V, Schmitz D, Sigrist SJ, Rosenmund C. 2016. RIM-binding protein 2 regulates release probability by fine-tuning calcium channel localization at murine hippocampal synapses. Proc Natl Acad Sci U S A 113(41):11615-11620.
Groc L, Lafourcade M, Heine M, Renner M, Racine V, Sibarita JB, Lounis B, Choquet D, Cognet L. 2007. Surface trafficking of neurotransmitter receptor: comparison between single-molecule/quantum dot strategies. J Neurosci 27(46):12433-12437.
Harris KM, Weinberg RJ. 2012. Ultrastructure of synapses in the mammalian brain. Cold Spring Harb Perspect Biol 4(5).
Heller JP, Michaluk P, Sugao K, Rusakov DA. 2017. Probing nano-organization of astroglia with multi-color super-resolution microscopy. J Neurosci Res.
Henkel AW, Simpson LL, Ridge RM, Betz WJ. 1996. Synaptic vesicle movements monitored by fluorescence recovery after photobleaching in nerve terminals stained with FM1-43. J Neurosci 16(12):3960-3967.
Hosy E, Butler C, Sibarita JB. 2014. Organization and dynamics of AMPA receptors inside synapses-nano-organization of AMPA receptors and main synaptic scaffolding proteins revealed by super-resolution imaging. Curr Opin Chem Biol 20:120-126.
Hoze N, Nair D, Hosy E, Sieben C, Manley S, Herrmann A, Sibarita JB, Choquet D, Holcman D. 2012. Heterogeneity of AMPA receptor trafficking and molecular interactions revealed by superresolution analysis of live cell imaging. Proc Natl Acad Sci U S A 109(42):17052-17057.
Hruska M, Henderson NT, Xia NL, Le Marchand SJ, Dalva MB. 2015. Anchoring and synaptic stability of PSD-95 is driven by ephrin-B3. Nat Neurosci 18(11):1594-1605.
Hua Y, Sinha R, Thiel CS, Schmidt R, Huve J, Martens H, Hell SW, Egner A, Klingauf J. 2011. A readily retrievable pool of synaptic vesicles. Nat Neurosci 14(7):833-839.
Huang B, Wang W, Bates M, Zhuang X. 2008. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319(5864):810-813.
Huang F, Sirinakis G, Allgeyer ES, Schroeder LK, Duim WC, Kromann EB, Phan T, Rivera-Molina FE, Myers JR, Irnov I, Lessard M, Zhang Y, Handel MA, Jacobs-Wagner C, Lusk CP, Rothman JE, Toomre D, Booth MJ, Bewersdorf J. 2016. Ultra-High Resolution 3D Imaging of Whole Cells. Cell 166(4):1028-1040.
Izeddin I, Specht CG, Lelek M, Darzacq X, Triller A, Zimmer C, Dahan M. 2011. Super-resolution dynamic imaging of dendritic spines using a low-affinity photoconvertible actin probe. PLoS One 6(1):e15611.
Ji N, Sato TR, Betzig E. 2012. Characterization and adaptive optical correction of aberrations during in vivo imaging in the mouse cortex. Proc Natl Acad Sci U S A 109(1):22-27.
Jones SA, Shim SH, He J, Zhuang X. 2011. Fast, three-dimensional super-resolution imaging of live cells. Nat Methods 8(6):499-508.
Juette MF, Gould TJ, Lessard MD, Mlodzianoski MJ, Nagpure BS, Bennett BT, Hess ST, Bewersdorf J. 2008. Three-dimensional sub-100 nm resolution fluorescence microscopy of thick samples. Nat Methods 5(6):527-529.
Jungmann R, Avendano MS, Dai M, Woehrstein JB, Agasti SS, Feiger Z, Rodal A, Yin P. 2016. Quantitative super-resolution imaging with qPAINT. Nat Methods 13(5):439-442.
Kamin D, Lauterbach MA, Westphal V, Keller J, Schonle A, Hell SW, Rizzoli SO. 2010. High- and low-mobility stages in the synaptic vesicle cycle. Biophys J 99(2):675-684.
Kellner RR, Baier CJ, Willig KI, Hell SW, Barrantes FJ. 2007. Nanoscale organization of nicotinic acetylcholine receptors revealed by stimulated emission depletion microscopy. Neuroscience 144(1):135-143.
Kiuchi T, Higuchi M, Takamura A, Maruoka M, Watanabe N. 2015. Multitarget super-resolution microscopy with high-density labeling by exchangeable probes. Nat Methods 12(8):743-746.
Klar TA, Jakobs S, Dyba M, Egner A, Hell SW. 2000. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc Natl Acad Sci U S A 97(15):8206-8210.
Lehmann M, Gottschalk B, Puchkov D, Schmieder P, Schwagerus S, Hackenberger CP, Haucke V, Schmoranzer J. 2015. Multicolor Caged dSTORM Resolves the Ultrastructure of Synaptic Vesicles in the Brain. Angew Chem Int Ed Engl 54(45):13230-13235.
Li TP, Song Y, MacGillavry HD, Blanpied TA, Raghavachari S. 2016. Protein Crowding within the Postsynaptic Density Can Impede the Escape of Membrane Proteins. J Neurosci 36(15):4276-4295.
Liu KS, Siebert M, Mertel S, Knoche E, Wegener S, Wichmann C, Matkovic T, Muhammad K, Depner H, Mettke C, Buckers J, Hell SW, Muller M, Davis GW, Schmitz D, Sigrist SJ. 2011. RIM-binding protein, a central part of the active zone, is essential for neurotransmitter release. Science 334(6062):1565-1569.
Lu HE, MacGillavry HD, Frost NA, Blanpied TA. 2014. Multiple spatial and kinetic subpopulations of CaMKII in spines and dendrites as resolved by single-molecule tracking PALM. J Neurosci 34(22):7600-7610.
MacGillavry HD, Hoogenraad CC. 2015. The internal architecture of dendritic spines revealed by super-resolution imaging: What did we learn so far? Exp Cell Res 335(2):180-186.
MacGillavry HD, Kerr JM, Blanpied TA. 2011. Lateral organization of the postsynaptic density. Mol Cell Neurosci 48(4):321-331.
MacGillavry HD, Kerr JM, Kassner J, Frost NA, Blanpied TA. 2016. Shank-cortactin interactions control actin dynamics to maintain flexibility of neuronal spines and synapses. Eur J Neurosci 43(2):179-193.
MacGillavry HD, Song Y, Raghavachari S, Blanpied TA. 2013. Nanoscale scaffolding domains within the postsynaptic density concentrate synaptic AMPA receptors. Neuron 78(4):615-622.
Maglione M, Sigrist SJ. 2013. Seeing the forest tree by tree: super-resolution light microscopy meets the neurosciences. Nat Neurosci 16(7):790-797.
Maidorn M, Rizzoli SO, Opazo F. 2016. Tools and limitations to study the molecular composition of synapses by fluorescence microscopy. Biochem J 473(20):3385-3399.
Manley S, Gillette JM, Patterson GH, Shroff H, Hess HF, Betzig E, Lippincott-Schwartz J. 2008. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat Methods 5(2):155-157.
Maschi D, Klyachko VA. 2017. Spatiotemporal Regulation of Synaptic Vesicle Fusion Sites in Central Synapses. Neuron.
Medintz IL, Uyeda HT, Goldman ER, Mattoussi H. 2005. Quantum dot bioconjugates for imaging, labelling and sensing. Nat Mater 4(6):435-446.
Metcalf DJ, Edwards R, Kumarswami N, Knight AE. 2013. Test Samples for Optimizing STORM Super-Resolution Microscopy. Jove-J Vis Exp(79).
Milovanovic D, Jahn R. 2015. Organization and dynamics of SNARE proteins in the presynaptic membrane. Front Physiol 6:89.
Minoshima M, Kikuchi K. 2017. Photostable and photoswitching fluorescent dyes for super-resolution imaging. J Biol Inorg Chem.
Mlodzianoski MJ, Juette MF, Beane GL, Bewersdorf J. 2009. Experimental characterization of 3D localization techniques for particle-tracking and super-resolution microscopy. Opt Express 17(10):8264-8277.
Nagerl UV, Willig KI, Hein B, Hell SW, Bonhoeffer T. 2008. Live-cell imaging of dendritic spines by STED microscopy. Proc Natl Acad Sci U S A 105(48):18982-18987.
Nair D, Hosy E, Petersen JD, Constals A, Giannone G, Choquet D, Sibarita JB. 2013. Super-resolution imaging reveals that AMPA receptors inside synapses are dynamically organized in nanodomains regulated by PSD95. J Neurosci 33(32):13204-13224.
Newpher TM, Ehlers MD. 2008. Glutamate receptor dynamics in dendritic microdomains. Neuron 58(4):472-497.
Nishimune H, Badawi Y, Mori S, Shigemoto K. 2016. Dual-color STED microscopy reveals a sandwich structure of Bassoon and Piccolo in active zones of adult and aged mice. Sci Rep 6:27935.
Owen DM, Williamson D, Magenau A, Gaus K. 2012. Optical techniques for imaging membrane domains in live cells (live-cell palm of protein clustering). Methods Enzymol 504:221-235.
Palay SL, Palade GE. 1955. The fine structure of neurons. J Biophys Biochem Cytol 1(1):69-88.
Panatier A, Arizono M, Nagerl UV. 2014. Dissecting tripartite synapses with STED microscopy. Philos Trans R Soc Lond B Biol Sci 369(1654):20130597.
Patton BR, Burke D, Owald D, Gould TJ, Bewersdorf J, Booth MJ. 2016. Three-dimensional STED microscopy of aberrating tissue using dual adaptive optics. Opt Express 24(8):8862-8876.
Pennacchietti F, Vascon S, Nieus T, Rosillo C, Das S, Tyagarajan SK, Diaspro A, Del Bue A, Petrini EM, Barberis A, Cella Zanacchi F. 2017. Nanoscale Molecular Reorganization of the Inhibitory Postsynaptic Density Is a Determinant of GABAergic Synaptic Potentiation. J Neurosci 37(7):1747-1756.
Perestenko PV, Henley JM. 2006. Visualization of AMPAR Trafficking and Surface Expression. In: Kittler JT, Moss SJ, editors. The Dynamic Synapse: Molecular Methods in Ionotropic Receptor Biology. Boca Raton (FL).
Perez de Arce K, Schrod N, Metzbower SW, Allgeyer E, Kong GK, Tang AH, Krupp AJ, Stein V, Liu X, Bewersdorf J, Blanpied TA, Lucic V, Biederer T. 2015. Topographic Mapping of the Synaptic Cleft into Adhesive Nanodomains. Neuron 88(6):1165-1172.
Petrini EM, Ravasenga T, Hausrat TJ, Iurilli G, Olcese U, Racine V, Sibarita JB, Jacob TC, Moss SJ, Benfenati F, Medini P, Kneussel M, Barberis A. 2014. Synaptic recruitment of gephyrin regulates surface GABAA receptor dynamics for the expression of inhibitory LTP. Nat Commun 5:3921.
Pleiner T, Bates M, Trakhanov S, Lee CT, Schliep JE, Chug H, Bohning M, Stark H, Urlaub H, Gorlich D. 2015. Nanobodies: site-specific labeling for super-resolution imaging, rapid epitope-mapping and native protein complex isolation. Elife 4:e11349.
Pougnet JT, Compans B, Martinez A, Choquet D, Hosy E, Boue-Grabot E. 2016. P2X-mediated AMPA receptor internalization and synaptic depression is controlled by two CaMKII phosphorylation sites on GluA1 in hippocampal neurons. Sci Rep 6:31836.
Pougnet JT, Toulme E, Martinez A, Choquet D, Hosy E, Boue-Grabot E. 2014. ATP P2X receptors downregulate AMPA receptor trafficking and postsynaptic efficacy in hippocampal neurons. Neuron 83(2):417-430.
Rizzoli SO. 2014. Synaptic vesicle recycling: steps and principles. EMBO J 33(8):788-822.
Rossier O, Octeau V, Sibarita JB, Leduc C, Tessier B, Nair D, Gatterdam V, Destaing O, Albiges-Rizo C, Tampe R, Cognet L, Choquet D, Lounis B, Giannone G. 2012. Integrins beta1 and beta3 exhibit distinct dynamic nanoscale organizations inside focal adhesions. Nat Cell Biol 14(10):1057-1067.
Rueckel M, Mack-Bucher JA, Denk W. 2006. Adaptive wavefront correction in two-photon microscopy using coherence-gated wavefront sensing. Proc Natl Acad Sci U S A 103(46):17137-17142.
Rusakov DA, Savtchenko LP, Zheng K, Henley JM. 2011. Shaping the synaptic signal: molecular mobility inside and outside the cleft. Trends Neurosci 34(7):359-369.
Rust MJ, Bates M, Zhuang X. 2006. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat Methods 3(10):793-795.
Salvatico C, Specht CG, Triller A. 2015. Synaptic receptor dynamics: from theoretical concepts to deep quantification and chemistry in cellulo. Neuropharmacology 88:2-9.
Schedin-Weiss S, Caesar I, Winblad B, Blom H, Tjernberg LO. 2016. Super-resolution microscopy reveals gamma-secretase at both sides of the neuronal synapse. Acta Neuropathol Commun 4:29.
Schoen M, Reichel JM, Demestre M, Putz S, Deshpande D, Proepper C, Liebau S, Schmeisser MJ, Ludolph AC, Michaelis J, Boeckers TM. 2015. Super-Resolution Microscopy Reveals Presynaptic Localization of the ALS/FTD Related Protein FUS in Hippocampal Neurons. Front Cell Neurosci 9:496.
Shim SH, Xia C, Zhong G, Babcock HP, Vaughan JC, Huang B, Wang X, Xu C, Bi GQ, Zhuang X. 2012. Super-resolution fluorescence imaging of organelles in live cells with photoswitchable membrane probes. Proc Natl Acad Sci U S A 109(35):13978-13983.
Shrivastava AN, Rodriguez PC, Triller A, Renner M. 2013. Dynamic micro-organization of P2X7 receptors revealed by PALM based single particle tracking. Front Cell Neurosci 7:232.
Sibarita JB. 2014. High-density single-particle tracking: quantifying molecule organization and dynamics at the nanoscale. Histochem Cell Biol 141(6):587-595.
Sidenstein SC, D'Este E, Bohm MJ, Danzl JG, Belov VN, Hell SW. 2016. Multicolour Multilevel STED nanoscopy of Actin/Spectrin Organization at Synapses. Sci Rep 6:26725.
Sieber JJ, Willig KI, Kutzner C, Gerding-Reimers C, Harke B, Donnert G, Rammner B, Eggeling C, Hell SW, Grubmuller H, Lang T. 2007. Anatomy and dynamics of a supramolecular membrane protein cluster. Science 317(5841):1072-1076.
Sigal YM, Speer CM, Babcock HP, Zhuang X. 2015. Mapping Synaptic Input Fields of Neurons with Super-Resolution Imaging. Cell 163(2):493-505.
Sinnen BL, Bowen AB, Forte JS, Hiester BG, Crosby KC, Gibson ES, Dell'Acqua ML, Kennedy MJ. 2017. Optogenetic Control of Synaptic Composition and Function. Neuron 93(3):646-660 e645.
Siskova Z, Justus D, Kaneko H, Friedrichs D, Henneberg N, Beutel T, Pitsch J, Schoch S, Becker A, von der Kammer H, Remy S. 2014. Dendritic structural degeneration is functionally linked to cellular hyperexcitability in a mouse model of Alzheimer's disease. Neuron 84(5):1023-1033.
Smith KR, Kopeikina KJ, Fawcett-Patel JM, Leaderbrand K, Gao R, Schurmann B, Myczek K, Radulovic J, Swanson GT, Penzes P. 2014. Psychiatric risk factor ANK3/ankyrin-G nanodomains regulate the structure and function of glutamatergic synapses. Neuron 84(2):399-415.
Specht CG, Izeddin I, Rodriguez PC, El Beheiry M, Rostaing P, Darzacq X, Dahan M, Triller A. 2013. Quantitative nanoscopy of inhibitory synapses: counting gephyrin molecules and receptor binding sites. Neuron 79(2):308-321.
Sudhof TC. 2006. Synaptic vesicles: an organelle comes of age. Cell 127(4):671-673.
Sydor AM, Czymmek KJ, Puchner EM, Mennella V. 2015. Super-Resolution Microscopy: From Single Molecules to Supramolecular Assemblies. Trends Cell Biol 25(12):730-748.
Szepesi Z, Hosy E, Ruszczycki B, Bijata M, Pyskaty M, Bikbaev A, Heine M, Choquet D, Kaczmarek L, Wlodarczyk J. 2014. Synaptically released matrix metalloproteinase activity in control of structural plasticity and the cell surface distribution of GluA1-AMPA receptors. PLoS One 9(5):e98274.
Takamori S, Holt M, Stenius K, Lemke EA, Gronborg M, Riedel D, Urlaub H, Schenck S, Brugger B, Ringler P, Muller SA, Rammner B, Grater F, Hub JS, De Groot BL, Mieskes G, Moriyama Y, Klingauf J, Grubmuller H, Heuser J, Wieland F, Jahn R. 2006. Molecular anatomy of a trafficking organelle. Cell 127(4):831-846.
Tang AH, Chen H, Li TP, Metzbower SR, MacGillavry HD, Blanpied TA. 2016. A trans-synaptic nanocolumn aligns neurotransmitter release to receptors. Nature 536(7615):210-214.
Tatavarty V, Das S, Yu J. 2012. Polarization of actin cytoskeleton is reduced in dendritic protrusions during early spine development in hippocampal neuron. Mol Biol Cell 23(16):3167-3177.
Tatavarty V, Kim EJ, Rodionov V, Yu J. 2009. Investigating sub-spine actin dynamics in rat hippocampal neurons with super-resolution optical imaging. PLoS One 4(11):e7724.
Teng KW, Ishitsuka Y, Ren P, Youn Y, Deng X, Ge P, Belmont AS, Selvin PR. 2016. Labeling proteins inside living cells using external fluorophores for microscopy. Elife 5.
Teng KW, Ishitsuka Y, Ren P, Youn Y, Deng X, Ge P, Lee SH, Belmont AS, Selvin PR. 2017. Labeling Proteins Inside Living Cells Using External Fluorophores for Fluorescence Microscopy. Elife 6.
Tillberg PW, Chen F, Piatkevich KD, Zhao Y, Yu CC, English BP, Gao L, Martorell A, Suk HJ, Yoshida F, DeGennaro EM, Roossien DH, Gong G, Seneviratne U, Tannenbaum SR, Desimone R, Cai D, Boyden ES. 2016. Protein-retention expansion microscopy of cells and tissues labeled using standard fluorescent proteins and antibodies. Nat Biotechnol 34(9):987-992.
Tonnesen J, Katona G, Rozsa B, Nagerl UV. 2014. Spine neck plasticity regulates compartmentalization of synapses. Nat Neurosci 17(5):678-685.
Tonnesen J, Nadrigny F, Willig KI, Wedlich-Soldner R, Nagerl UV. 2011. Two-color STED microscopy of living synapses using a single laser-beam pair. Biophys J 101(10):2545-2552.
Turkowyd B, Virant D, Endesfelder U. 2016. From single molecules to life: microscopy at the nanoscale. Anal Bioanal Chem 408(25):6885-6911.
Urban NT, Willig KI, Hell SW, Nagerl UV. 2011. STED nanoscopy of actin dynamics in synapses deep inside living brain slices. Biophys J 101(5):1277-1284.
Varela JA, Ferreira JS, Dupuis JP, Durand P, Bouchet D, Groc L. 2016. Single nanoparticle tracking of [Formula: see text]-methyl-d-aspartate receptors in cultured and intact brain tissue. Neurophotonics 3(4):041808.
Voigt A, Freund R, Heck J, Missler M, Obermair GJ, Thomas U, Heine M. 2016. Dynamic association of calcium channel subunits at the cellular membrane. Neurophotonics 3(4):041809.
Wang K, Milkie DE, Saxena A, Engerer P, Misgeld T, Bronner ME, Mumm J, Betzig E. 2014. Rapid adaptive optical recovery of optimal resolution over large volumes. Nat Methods 11(6):625-628.
Wang L, Dumoulin A, Renner M, Triller A, Specht CG. 2016. The Role of Synaptopodin in Membrane Protein Diffusion in the Dendritic Spine Neck. PLoS One 11(2):e0148310.
Wen PJ, Grenklo S, Arpino G, Tan X, Liao HS, Heureaux J, Peng SY, Chiang HC, Hamid E, Zhao WD, Shin W, Nareoja T, Evergren E, Jin Y, Karlsson R, Ebert SN, Jin A, Liu AP, Shupliakov O, Wu LG. 2016. Actin dynamics provides membrane tension to merge fusing vesicles into the plasma membrane. Nat Commun 7:12604.
Westphal V, Rizzoli SO, Lauterbach MA, Kamin D, Jahn R, Hell SW. 2008. Video-rate far-field optical nanoscopy dissects synaptic vesicle movement. Science 320(5873):246-249.
Wijetunge LS, Angibaud J, Frick A, Kind PC, Nagerl UV. 2014. Stimulated emission depletion (STED) microscopy reveals nanoscale defects in the developmental trajectory of dendritic spine morphogenesis in a mouse model of fragile X syndrome. J Neurosci 34(18):6405-6412.
Wilhelm BG, Mandad S, Truckenbrodt S, Krohnert K, Schafer C, Rammner B, Koo SJ, Classen GA, Krauss M, Haucke V, Urlaub H, Rizzoli SO. 2014. Composition of isolated synaptic boutons reveals the amounts of vesicle trafficking proteins. Science 344(6187):1023-1028.
Willig KI, Barrantes FJ. 2014. Recent applications of superresolution microscopy in neurobiology. Curr Opin Chem Biol 20:16-21.
Willig KI, Rizzoli SO, Westphal V, Jahn R, Hell SW. 2006. STED microscopy reveals that synaptotagmin remains clustered after synaptic vesicle exocytosis. Nature 440(7086):935-939.
Willig KI, Steffens H, Gregor C, Herholt A, Rossner MJ, Hell SW. 2014. Nanoscopy of filamentous actin in cortical dendrites of a living mouse. Biophys J 106(1):L01-03.
Yamanaka M, Smith NI, Fujita K. 2014. Introduction to super-resolution microscopy. Microscopy (Oxf) 63(3):177-192.
Younts TJ, Monday HR, Dudok B, Klein ME, Jordan BA, Katona I, Castillo PE. 2016. Presynaptic Protein Synthesis Is Required for Long-Term Plasticity of GABA Release. Neuron 92(2):479-492.
Zhang J, Carver CM, Choveau FS, Shapiro MS. 2016. Clustering and Functional Coupling of Diverse Ion Channels and Signaling Proteins Revealed by Super-resolution STORM Microscopy in Neurons. Neuron 92(2):461-478.
Zhong H. 2015. Applying superresolution localization-based microscopy to neurons. Synapse.